Sidebar
 This version (2018/06/20 13:12) is a draft.
This version (2018/06/20 13:12) is a draft.Approvals: 0/1
This is an old revision of the document!
sxviper
Initial 3D Model - VIPER: ab initio 3D structure determination using Validation of Individual Parameter Reproducibility (VIPER). Designed to determine a validated initial model using a small set of class averages produced by ISAC2.
Usage
Usage in command line
sxviper.py stack directory --radius=outer_radius --sym=sym --ir=inner_radius --rs=ring_step --xr=x_range --yr=y_range --ts=translational_search_step --delta=angular_step --center=center_type --maxit1=max_iter1 --maxit2=max_iter2 --mask3D=mask3D --moon_elimination=moon_elimination --L2threshold=L2threshold --ref_a=ref_a --nruns=nruns --doga=doga --fl=fl --aa=aa --pwreference=pwreference --debug
Typical usage
sxrviper exists only in MPI version.
mpirun --npernode 16 -np 24 --host node1,node2 sxviper.py stack output_directory --fl=0.25 --radius=30 --xr=2 --moon_elimination=750,4.84
A fast track option, that can be used to choose parameters in the appropriate ranges (for example, obtaining adequate spatial frequency filtering --fl) is provided below. Since it employs extreme values for some parameters this command can be used only for parameter tuning for the VIPER algorithm.
mpirun --npernode 16 -np 16 --host node1 sxviper.py stack output_directory --fl=0.25 --radius=30 --xr=1 --nruns=2 --L2threshold=1.0e300 --doga=-1
The VIPER program needs MPI environment to work properly. Number of used MPI processes must be a multiple of --nruns (default = 6).
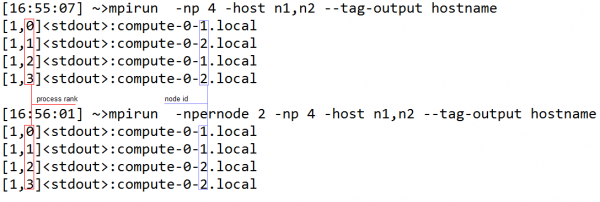
Since VIPER makes use of group of processors working together, it is important from a time efficiency point of view to have processors within a group being allocated on the same node. This way any data exchange within the group does not use network traffic. The --npernode option of mpirun is useful in accomplishing this goal. As shown in the example below when --npernode is used mpi allocates the ranks of the processors sequentially, not moving to the next node until the current one is filled. If --npernode is not used then processors are allocated in a round robin fashion (i.e. jumping to the next node with each allocation). Since in VIPER, groups contain consecutively ranked processors, it is important to provide “--npernode XX”, where XX is the number of processors per node.
Input
Main Parameters
- stack
- Input images stack: A small set of class averages produced by ISAC2. (default required string)
- directory
- Output directory: The directory will be automatically created and the results will be written here. If the directory already exists, results will be written there, possibly overwriting previous runs. (default required string)
- --radius
- Target particle radius [Pixels]: Use the same value as in ISAC2. It has to be less than half the box size. (default 29)
- --sym
- Point-group symmetry: Point-group symmetry of the target particle. (default c1)
- --moon_elimination
- Eliminate disconnected regions: Used to removed disconnected pieces from the model. It requires as argument a comma separated string with the mass in KDa and the pixel size. (default none)
Advanced Parameters
- --ir
- Inner rotational search radius [Pixels]: Inner rotational search radius [Pixels]. (default 1)
- --rs
- Ring step size [Pixels]: Step between rings used for the rotational search. (default 1)
- --xr
- X search range [Pixels]: The translational search range in the x direction will take place in a +/xr range. (default '0')
- --yr
- Y search range [Pixels]: The translational search range in the y direction. If omitted it will be xr. (default '0')
- --ts
- Translational search step [Pixels]: The search will be performed in -xr, -xr+ts, 0, xr-ts, xr, can be fractional. (default '1.0')
- --delta
- Projection angular step [Degrees]: Projection angular step. (default '2.0')
- --center
- Center 3D template: 0: no centering; 1: center of gravity (default -1.0)
- --maxit1
- Maximum iterations - GA step: Maximum iterations for GA step. (default 400)
- --maxit2
- Maximum iterations - Finish step: Maximum iterations for Finish step. (default 50)
- --mask3D
- 3D mask: Path to 3D mask file. (default sphere)
- --L2threshold
- GA stop threshold: Defines the maximum relative dispersion of volumes' L2 norms. (default 0.03)
- --ref_a
- Projection generation method: Method for generating the quasi-uniformly distributed projection directions. S - Saff algorithm, or P - Penczek 1994 algorithm. (default S)
- --nruns
- GA population size: This defines the number of quasi-independent volumes generated. (default 6)
- --doga
- Threshold to start GA: Do GA when the fraction of orientation that changes less than 1.0 degrees is at least this fraction. (default 0.1)
- --fl
- Low-pass filter frequency [1/Pixels]: Using a hyperbolic tangent low-pass filter. Specify with absolute frequency. (default 0.25)
- --aa
- Low-pass filter fall-off [1/Pixels]: Fall-off of for the hyperbolic tangent low-pass filter. Specify with absolute frequency. (default 0.1)
- --pwreference
- Power spectrum reference: Text file containing a 1D reference power spectrum. (default none)
- --debug
- Verbose: Print debug info. (default False)
Output
Description
- This program uses a user-defined projection angle and translation shift to perform 3D reconstruction. The translation shifts, and step are not limited to integer number. For a given delta, the program will perform maxit round refinement. So the final refinement iteration is maxit*(number of delta values).
- For the program to work, attributes xform.projection (Transform object containing three Euler angles and two in-plane shifts) have to be set in the header of each file. If their values are not known, all should be set to zero.
- The program will start alignment from the current alignment parameters xform.projection stored in file headers.
- The program only change the alignment parameters in their header. The images in stack keep untouched. (Neither rotated nor shifted.)
Method
Reference
Developer Notes
Author / Maintainer
Pawel A. Penczek
Keywords
Category 1:: APPLICATIONS
Files
sparx/bin/sxviper.py
See also
Maturity
Beta:: Under evaluation and testing. Please let us know if there are any bugs.
Bugs
There are no known bugs so far.