Sidebar
Approvals: 0/1
Table of Contents
crYOLO 1.1.0 release notes
Changes crYOLO
- crYOLO now supports filaments (more here)
- New evaluation tool (more here)
- Supports empty box files for training on particle-free images
- Extended data augmentation: Horizontal flip and flip along both axes
- Experimental support of periodic restarts during training (with –warm_restarts)
Changes crYOLO Boxmanager
- Support of visualization of EMAN2 helical coordinates (particle coordinates)
- New boxes could be loaded with a new color while keeping the old.
- Several bug fixes
Picking filaments - Using a model trained for your data
Since version 1.1.0 crYOLO supports picking filaments.
Data preparation
As described previously, filtering your image using a low-pass filter is probably a good idea.
After this is done, you have to prepare training data for your model.
 Right now, you have to use the e2helixboxer.py to generate the training data:
Right now, you have to use the e2helixboxer.py to generate the training data:
e2helixboxer.py --gui my_images/*.mrc
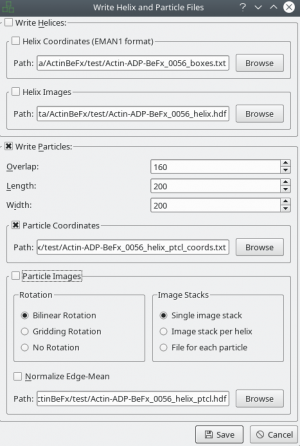
After tracing your training data in e2helixboxer, export them using File → Save. Make sure that you export particle coordinates as this the only format supported right now (see screenshot). In the following example, it is expected that you exported into a folder called “train_annot”.
Configuration
You can configure it the same way as for a “normal” project. We recommend to use patch mode.
Training
In principle, there is not much difference in training crYOLO for filament picking and particle picking. For project with roughly 20 filaments per image we successfully trained on 40 images (⇒ 800 filaments). However, in our experience the warm-up phase and training need a little bit more time:
1. Warm up your network
cryolo_train.py -c config.json -w 10
2. Train your network
cryolo_train.py -c config.json -w 0 -e 10
The final model will be called model.h5
Picking
The biggest difference in picking filaments with crYOLO is during prediction. However, there are just three additional parameters needed:
- - -filament: Option that tells crYOLO that you want to predict filaments
- -fw: Filament width (pixels)
- -bd: Inter-Box distance (pixels).
Let's assume you want to pick a filament with a width of 100 pixels. The box size is 200×200 and you want a 90% overlap, than the picking command would be:
cryolo_predict.py -c config.json -w model.h5 --filament -fw 100 -bd 20 -o boxes/ -g 0
The directory boxes will be created and all results are saved there. The format is the eman2 helix format with particle coordinates.
Visualize the results
You can use the boxmanager as described previously.
Evaluate your results
The new evaluation tool allows you, based on your validation data, to get statistics about your training. If you followed the tutorial, the validation data are selected randomly. With crYOLO 1.1.0 a run file for each training is created and saved into the folder runfiles/ in your project directory. This run file contains which files were selected for validation, and you can run your evaluation as follows:
cryolo_evaluation.py -c config.json -w model.h5 -r runfiles/run_20180821-144617.json
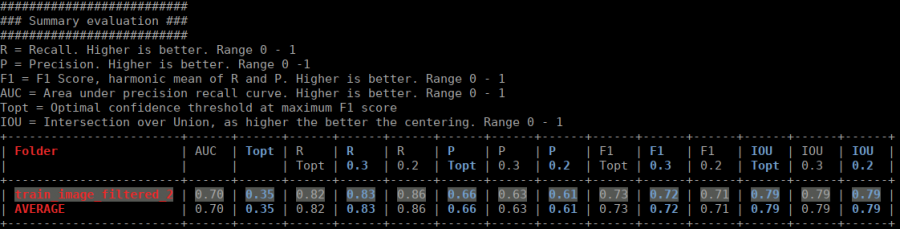
The table contains several statistics:
- AUC: Area under curve of the precision-recall curve. Overall summary statistics. Perfect classifier = 1, Worst classifier = 0
- Topt: Optimal confidence threshold with respect to the F1 score. It might not be ideal for your picking, as the F1 score weighs recall and precision equally. However in SPA, recall is often more important than the precision.
- R (Topt): Recall using the optimal confidence threshold.
- R (0.3): Recall using a confidence threshold of 0.3.
- R (0.2): Recall using a confidence threshold of 0.2.
- P (Topt): Precision using the optimal confidence threshold.
- P (0.3): Precision using a confidence threshold of 0.3.
- P (0.2): Precision using a confidence threshold of 0.2.
- F1 (Topt): Harmonic mean of precision and recall using the optimal confidence threshold.
- F1 (0.3): Harmonic mean of precision and recall using a confidence threshold of 0.3.
- F1 (0.2): Harmonic mean of precision and recall using a confidence threshold of 0.2.
- IOU (Topt): Intersection over union of the auto-picked particles and the corresponding ground-truth boxes. The higher, the better – evaluated with the optimal confidence threshold.
- IOU (0.3): Intersection over union of the auto-picked particles and the corresponding ground-truth boxes. The higher, the better – evaluated with a confidence threshold of 0.3.
- IOU (0.2): Intersection over union of the auto-picked particles and the corresponding ground-truth boxes. The higher, the better – evaluated with a confidence threshold of 0.2.
If the training data consists of multiple folders, then evaluation will be done for each folder separately.
General model
Increase the number of hand picked datasets to 25 by adding:
- Add EMPIAR 10154 (Thanks to Daniel Prumbaum)
- Add EMPIAR 10186 (Thanks to Sebastian Tacke)
- Add EMPIAR 10097 Hemagglutinin (Thanks to Birte Siebolds)
- Add EMPIAR 10081 HCN1 (Thanks to Pascel Lill)
- Add internal dataset (Thanks to Daniel Roderer)
- Furthermore we added 8 datasets of protein-free grids (Thanks to Tobias Raisch and Daniel Prumbaum)