Sidebar
 This version is outdated by a newer approved version.
This version is outdated by a newer approved version. This version (2019/09/12 16:43) is a draft.
This version (2019/09/12 16:43) is a draft.Approvals: 0/1
This is an old revision of the document!
sp_separate_class
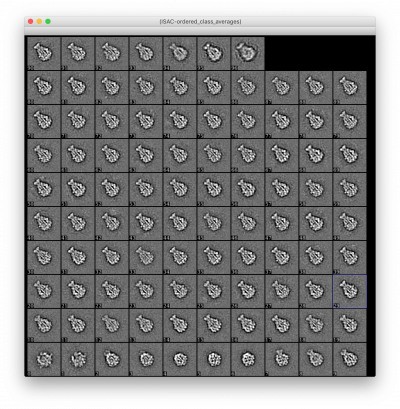
Separate Into Classes : Separates stacks of particle images into stacks for each class.
Usage
Usage in command line:
sp_separate_class.py input_class_avgs input_image_stack output_directory --filtrad=filter_radius --apix=pixel_size --shrink=shrink_factor --align_isac_dir=isac_directory --format=stack_format --verbose
Typical usage
The purpose of sp_separate_class.py is to:
: extract particle-membership information from a stack of class averages : write particle-membership lists for each class, and : write separate stacks for each class, with an option to low-pass filter and/or downsample the images
1. Standard usage: create separate stacks for each class:
sp_separate_class.py input_class_avgs input_image_stack output_directory
2. Apply a low-pass filter to the image stacks:
sp_separate_class.py input_class_avgs input_image_stack output_directory --filt=filter_radius --apix=pixel_size
Filter radius is in units of Angstroms. If apix parameter is not specified, program will assume units of pixels^-1.
3. Downsample output image stack:
sp_separate_class.py input_class_avgs input_image_stack output_directory --shrink=shrink_factor
4. Apply ISAC alignments to particles:
sp_separate_class.py input_class_avgs input_image_stack output_directory --align_isac_dir=isac_directory
If the input class averages are ordered_class_averages.hdf, the alignments applied to the ordered class averages will be applied to the particles.
Input
Main Parameters
- input_class_avgs
- Set of 2D class averages, with particle-membership information in header. (default required string)
- input_image_stack
- Particle image stack. (default required string)
- output_directory
- Directory where outputs will be written. (default required string)
- --filtrad
- Gaussian low-pass filter radius, Angstroms if apix specified below, else pixels^-1. (default None)
- --apix
- Angstroms per pixel, might be downsampled already by ISAC2. (default None)
- --shrink
- Downsampling factor, e.g., 6 → 1/6 original size. (default None)
- --align_isac_dir
- If applying alignments, directory for ISAC output (default None)
- --verbose
- Writes additional messages to the terminal during execution. (default False)
Advanced Parameters
- --format
- Format of optional output aligned-imaged stacks. (default .mrcs)
Output
- classmap.txt
- Class-to-particle lookup table, one file for all classes
- docclass???.txt
- List of particles for each class, one file per class
- params_combined.txt
- (Optional) Combined particle alignment parameters
- stack_all.bdb
- Virtual stack with all particles
- stkclass_???.bdb
- Virtual stacks of particles for each class
- stkclass_???.bdb
- Virtual stacks of unaligned particles for each class
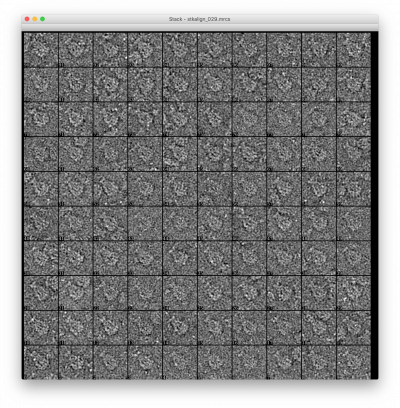
- stkalign_???.bdb
- (Optional) Virtual stacks of aligned and optionally filtered particles for each class
Description
Method
Reference
Developer Notes
: Should allow filter types other than Gaussian low-pass
Author / Maintainer
Tapu Shaikh
Keywords
Category 1:: APPLICATIONS
Files
sphire/bin/sp_separate_class.py
See also
Maturity
Beta:: Under evaluation and testing. Please let us know if there are any bugs.
Bugs
There are no known bugs so far.